陳亭妏副教授研究團隊發表研究成果於 Bioinformatics
連結網址:https://academic.oup.com/bioinformatics/article/38/18/4286/6649618
Abstract
Motivation: Microbiota analyses have important implications for health and science. These analyses make use of 16S/18S rRNA gene sequencing to identify taxa and predict species diversity. However, most available tools for analyzing microbiota data require adept programming skills and in-depth statistical knowledge for proper implementation. While long-read amplicon sequencing can lead to more accurate taxa predictions and is quickly becoming more common, practitioners have no easily accessible tools with which to perform their analyses.
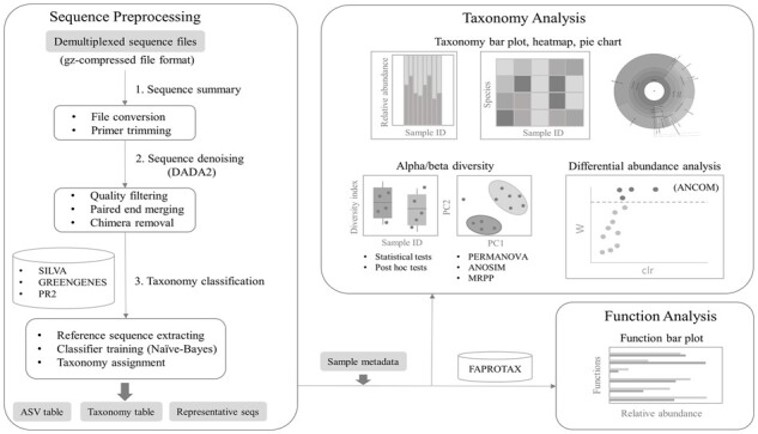
Results: We present MOCHI, a GUI tool for microbiota amplicon sequencing analysis. MOCHI preprocesses sequences, assigns taxonomy, identifies different abundant species and predicts species diversity and function. It takes either taxonomic count table or FASTQ of partial 16S/18S rRNA or full-length 16S rRNA gene as input. It performs analyses in real time and visualizes data in both tabular and graphical formats.
Availability and implementation: MOCHI can be installed to run locally or accessed as a web tool at https://mochi.life.nctu.edu.tw